27/03/2013
For decades, the concept of cell fate was largely defined by Conrad Hal Waddington's metaphor: a ball rolling irreversibly down a one-way slope, its path determined and progressively restricted as it differentiated. This view painted a picture of terminal differentiation, where a cell, once committed to a specific identity, could not turn back or change its lineage. However, a series of pivotal discoveries in cell biology have dramatically reshaped this traditional perspective, revealing an astonishing plasticity within somatic cells. This shift in understanding has not only deepened our knowledge of biological development but has also opened up unprecedented avenues for regenerative medicine, particularly through the powerful technique known as direct reprogramming.
Direct reprogramming, also referred to as transdifferentiation, is a revolutionary process that enables the conversion of one somatic cell type directly into another, bypassing the need to first revert to an intermediary pluripotent state, such as an induced pluripotent stem cell (iPSC). Unlike iPSC reprogramming, which involves a 'detour' through a stem cell phase before redifferentiating into the desired cell type, direct reprogramming offers a more streamlined and, in principle, faster route to cell fate conversion. This elegant simplicity holds immense promise for therapeutic applications, allowing for the potential generation of functional cells directly within damaged tissues, thereby avoiding complex ex vivo expansion and transplantation procedures.
- The Evolution of a Concept: From Fixed Fate to Flexible Identity
- Direct Reprogramming: A Promising Avenue for Regenerative Medicine
- Unravelling the Molecular Machinery of Cell Fate Conversion
- Advancing the Science: Enhancing Efficiency and Maturation
- Discovering New Reprogramming Factors: The Future of Cell Conversion
- Challenges and the Road Ahead for Clinical Application
The Evolution of a Concept: From Fixed Fate to Flexible Identity
The first crack in Waddington's seemingly immutable developmental landscape appeared in 1987 with the discovery that overexpressing MYOD, a transcription factor typically found in skeletal muscle cells, could convert mouse embryonic fibroblasts into myoblasts. This landmark finding demonstrated for the first time that cell identity was not as fixed as previously believed and could, in fact, be manipulated. In the decades that followed, a substantial body of research has built upon this initial revelation, showing that cell fate can be altered through the forced expression of various transcription factors, non-coding RNAs, and even through the application of small molecules.
Early successes in direct reprogramming often involved conversions between cells derived from the same embryonic germ layer. For instance, the expression of Gata1 was shown to convert avian myeloblasts into eosinophils or thromboblast-like cells. Similarly, the combination of Neurog3, Pdx1, and Mafa was found to reprogramme adult pancreatic exocrine cells into functional insulin-secreting cells in mice. However, the field truly broadened with the discovery of iPSCs in 2006, which inspired further exploration into converting somatic cells into entirely different lineages, even across germ layer barriers. By 2010, induced cardiomyocytes (iCMs) were successfully generated from mouse cardiac fibroblasts, and soon after, induced neurons (iNs) and induced hepatocytes followed. This demonstrated the remarkable versatility of direct reprogramming, proving that cells from one germ layer (e.g., mesoderm-derived fibroblasts) could be converted into cell types originating from another (e.g., ectoderm-derived neurons).
Direct Reprogramming: A Promising Avenue for Regenerative Medicine
Beyond its contributions to our fundamental understanding of cell fate specification and plasticity, direct reprogramming offers significant advantages for regenerative medicine. Compared to iPSC-based therapies, direct reprogramming is generally faster and more efficient. Crucially, its ability to convert cells in situ – directly within the target tissue – removes the need for laborious ex vivo cell expansion and subsequent transplantation. This 'on-site' conversion minimises the risk of immune rejection, simplifies delivery, and potentially reduces the complexities and costs associated with traditional cell therapies.
For example, in vivo direct reprogramming has successfully generated pancreatic β-cells in diabetic mouse models, leading to increased insulin levels and improved glucose tolerance. Similarly, scar-forming cardiac fibroblasts have been converted into iCMs in mice after myocardial infarction, resulting in reduced scar size and improved heart function. Endogenous glial cells have been transformed into functional neurons, hepatic myofibroblasts into hepatocytes to attenuate liver fibrosis, and rod photoreceptors generated in retinas, leading to improved vision. These successes highlight the profound therapeutic potential of direct reprogramming across various organ systems.
Comparison: Direct Reprogramming vs. iPSC Reprogramming
| Feature | Direct Reprogramming (Transdifferentiation) | iPSC Reprogramming |
|---|---|---|
| Intermediate State | No pluripotent intermediate | Requires pluripotent intermediate (iPSC) |
| Speed | Generally faster | Slower (requires pluripotency induction then differentiation) |
| Efficiency | Often higher for specific conversions | Can be high for pluripotency, but subsequent differentiation can be inefficient |
| Epigenetic Memory | May retain epigenetic hallmarks of original cell (e.g., ageing) | Erases most epigenetic memory during pluripotency |
| In Situ Application | Highly suitable for in vivo (on-site) conversion | Typically requires ex vivo expansion and transplantation |
| Tumorigenic Risk | Lower risk (no pluripotent state) | Potential for teratoma formation from residual iPSCs |
| Complexity | Potentially simpler delivery (e.g., direct injection) | Complex isolation, expansion, and differentiation protocols |
Unravelling the Molecular Machinery of Cell Fate Conversion
To truly harness the power of direct reprogramming for clinical applications, a deep understanding of its underlying molecular mechanisms is essential. Research has focused on elucidating the roles of transcription factors, epigenetic modifications, non-coding RNAs, and even metabolic shifts in orchestrating these remarkable cell fate changes.
The Orchestrators: Transcription Factors
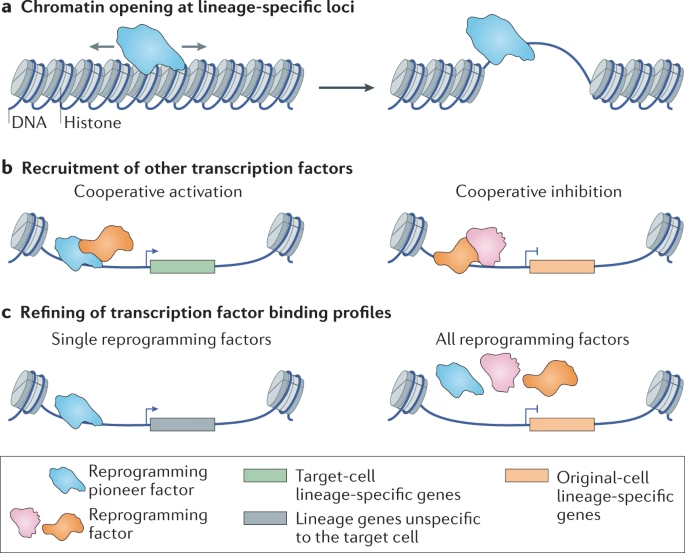
Transcription factors (TFs) are the molecular maestros of lineage conversion. They are crucial for dictating lineage-specific gene expression. A special class, known as pioneer factors, are particularly important. Unlike other TFs, pioneer factors can bind to and 'open' closed chromatin regions, making them accessible for other canonical TFs to bind and initiate gene expression. Examples include Gata4 in cardiac reprogramming and Ascl1 in neuronal reprogramming.
TFs often work cooperatively. For instance, Ascl1 recruits Brn2 to neuronal targets to induce cell fate conversion. In cardiac reprogramming, multiple TFs often co-occupy DNA binding sites, showing synergistic interactions that lead to stronger activation of heart-related gene programmes. Interestingly, TFs can also refine each other's binding profiles; the pattern of cardiac reprogramming factors expressed individually can differ significantly from when they are expressed together. The optimal dosage ratio of these factors is also critical, as some, like Mef2c in cardiac reprogramming, play a more dominant role in initial gene activation and subsequent cell maturation than others.
Epigenetic Remodelling: Shaping the Cell's Destiny
Epigenetic modifications, which involve changes to DNA or histone proteins that affect gene expression without altering the underlying DNA sequence, are central to direct reprogramming. These modifications act as a crucial layer of control, dictating which genes are active or silent within a cell.
Chromatin Accessibility: Opening Up New Possibilities
The forced expression of reprogramming factors induces drastic and rapid changes in chromatin accessibility – how tightly DNA is packed within the nucleus. Studies have shown that chromatin remodelling can occur as early as 12 hours after introducing reprogramming factors, primarily at regions far from the transcription start sites. Typically, regions that define the original cell type become less accessible, indicating transcriptional repression, while new regions associated with the target cell type become more accessible. For example, in cardiac reprogramming, regions related to heart and striated muscle development quickly become accessible. The cooperative interaction of reprogramming factors is believed to drive these changes, although the effect can be context-dependent, with some TFs like Ascl1 consistently increasing accessibility, while others can have varied effects.
Histone Modifications: The Epigenetic Code
Histone post-translational modifications are key regulators of gene expression. Several modifications play critical roles in direct reprogramming:
- H3K4me3 (Histone 3 Lysine 4 Trimethylation): Associated with active transcription, it marks promoters of active genes and is a hallmark of successful activation of the target cell's transcriptional programme. It's rapidly deposited at new lineage-specific genes and removed from old ones.
- H3K27me3 (Histone 3 Lysine 27 Trimethylation): A repressive mark, its removal from target genes and gain on original cell-specific genes is crucial for successful reprogramming. Inhibiting H3K27me3 methyltransferases can facilitate reprogramming.
- H3K9me3 (Histone 3 Lysine 9 Trimethylation): Another repressive mark associated with heterochromatin (tightly packed DNA) that typically prevents TF binding. It acts as a major barrier to reprogramming, and its temporary erasure can enhance efficiency in some contexts (e.g., cardiac), but not others (e.g., neuronal).
- H3K27ac (Histone 3 Lysine 27 Acetylation): Positively correlated with gene expression, it marks active enhancers and super-enhancers, which are crucial for establishing cell identity. Its presence often correlates with reprogramming factor binding.
- H2AK119Ub (Histone H2A Lysine 119 Ubiquitylation): This repressive mark, when present at target loci in the starting cells, can impede reprogramming. Its removal, for example by depleting Bmi1, has been shown to significantly enhance cardiac reprogramming efficiency.
Furthermore, multiple histone modifications can coexist on the same histone, cooperatively regulating transcription. Bivalent marks (e.g., H3K4me3 and H3K27me3) can keep lineage-specific genes in a silent but poised state, ready for rapid activation. Trivalent chromatin domains (e.g., H3K27ac, H3K4me1, H3K9me3) have also been identified as important for enabling pioneer factor binding and enhancing reprogramming efficiency in certain contexts.
DNA Methylation: Silencing the Past
DNA methylation involves the addition of methyl groups to DNA bases, typically leading to gene silencing. A global reconfiguration of DNA methylation patterns is observed during various direct reprogramming processes. For instance, promoters of cardiac lineage genes become demethylated early during iCM conversion, while fibroblast-specific genes undergo de novo methylation. This dynamic interplay ensures the silencing of the old cell identity and the activation of the new one, highlighting its crucial role in successful cell fate conversion.
Beyond Genes: Non-Coding RNAs and Metabolic Shifts
Beyond transcription factors and epigenetic modifications, other molecular players, such as non-coding RNAs and metabolic factors, are gaining recognition for their roles in direct reprogramming.
MicroRNAs (miRNAs) are small RNAs that regulate gene expression at the post-transcriptional level. Their small size makes them easier to deliver than DNA or mRNA encoding TFs, making them valuable tools to refine direct reprogramming. For example, the addition of miR-133a can improve cardiac reprogramming efficiency by silencing multiple downstream effectors. Similarly, miR-9/9* and miR-124 have been used in neuronal reprogramming, reconfiguring chromatin accessibility and DNA methylation, and disrupting repressive transcription factors like REST. Long non-coding RNAs (lncRNAs), which are longer than 200 nucleotides, also regulate gene expression by modulating epigenetics and 3D chromosome structure. While their role in direct reprogramming is still being explored, isoforms of lnc-NR2F1 have been found to have opposite effects on neuronal reprogramming, suggesting a delicate balance in controlling cell fate.

Metabolic reprogramming, though mentioned in the provided text as an area of recent substantial progress, is not detailed further. However, it's increasingly understood that changes in cellular metabolism are intimately linked with cell identity and can significantly influence the efficiency and outcome of cell fate conversions.
Advancing the Science: Enhancing Efficiency and Maturation
Despite the immense promise, direct reprogramming faces several challenges, including low conversion efficiency, the immaturity of reprogrammed cells, and the need for safe and precise delivery methods. Significant progress is being made to overcome these hurdles.
Identifying Optimal Starting Cells
The choice of starting cell is crucial. Ideally, these cells should be abundant and amenable to reprogramming. Macroglial cells like astrocytes and oligodendrocyte progenitor cells (NG2 glia) have been extensively explored for conversion to iNs, given their widespread distribution in the central nervous system and neurogenic potential. Cardiac fibroblasts are a major source for iCM conversion, especially after heart injury where they contribute to scar formation. In the pancreas, abundant exocrine acinar cells and easily convertible α-cells are considered ideal for generating β-cells. It's also important to note that the native cellular context can influence susceptibility to reprogramming; for example, astrocytes from different brain regions show varying amenability.
The Power of the Native Microenvironment
One of the most exciting findings is that in vivo reprogramming often yields cells that are more mature and functional than those obtained in vitro. This is attributed to the unique cellular and extracellular environment of live organs, which provides tissue-specific biochemical and mechanical signals. For instance, in vivo generated iCMs resemble adult cardiomyocytes in their transcriptional profile, structure, and physiology, becoming electrically coupled with endogenous cells. Similarly, β-cells generated in vivo are functionally comparable to native ones. The native microenvironment, including growth factors (like FGF2) and the biophysical properties of the extracellular matrix (e.g., 3D scaffolds, mechanical force), plays a crucial role in enhancing cell fate conversion and maturation.
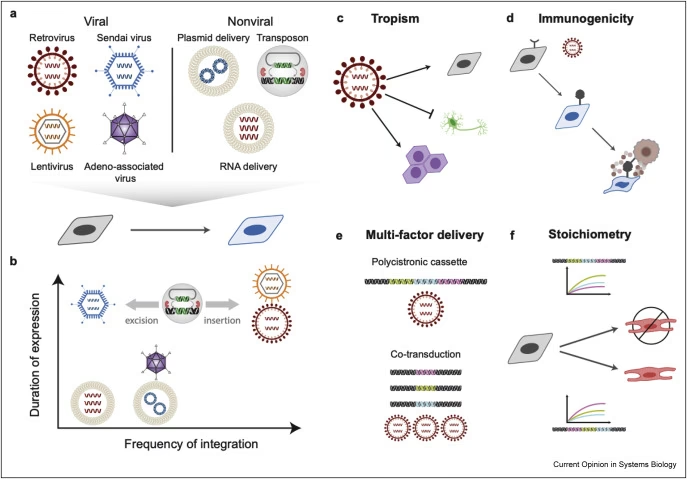
Refining Reprogramming Factor Delivery
Early studies often relied on viral vectors for in vivo delivery, which carried risks of genome integration and potential tumorigenesis. Significant progress has been made in developing safer, integration-free strategies, including Sendai virus, modified mRNA, single guide RNA (sgRNA), and nanoparticle-based gene carriers. Small molecules have emerged as a particularly promising alternative due to their non-immunogenic nature, cost-effectiveness, and ease of standardisation. While challenges remain in extending their duration of action and precisely controlling their function in vivo, advances in hydrogel technology are facilitating controlled drug release, allowing for sustained exposure to small molecules and improved reprogramming efficacy.
Discovering New Reprogramming Factors: The Future of Cell Conversion
With nearly 2,000 transcription factors in the mammalian genome, traditional exhaustive screening to identify reprogramming factors is a slow process. Researchers are now adopting innovative strategies to expedite discovery:
- Algorithm-based Prediction: Computational frameworks like Mogrify predict TF sets capable of converting one cell type to another based on transcriptomic information and interactome databases. This approach has successfully identified factors to convert fibroblasts to keratinocytes or keratinocytes to microvascular endothelial cells.
- CRISPR-activation Screening: This high-throughput method uses a catalytically dead Cas9 endonuclease fused to a transcription activator to activate endogenous gene expression. It allows for unbiased, genome-wide screens of TFs, leading to the identification of highly efficient reprogramming factors, such as endogenous Brn2 and Ngn1 for neuronal reprogramming, achieving significantly higher efficiencies than traditional forced expression methods.
These new strategies are crucial for uncovering novel reprogramming factors and optimising existing 'cocktails' to achieve even more efficient and precise cell fate conversions.
Challenges and the Road Ahead for Clinical Application
Despite the remarkable progress, several significant challenges must be overcome before direct reprogramming can be routinely translated into clinical therapies. The efficiency of conversion, while improving, can still be low for some cell types, and maintaining the full maturity and functional integration of reprogrammed cells remains a hurdle. Furthermore, ensuring the safety of delivery methods, particularly for in vivo applications, is paramount to prevent off-target effects or unintended cellular changes.
Another critical consideration is the variability in patient response. Studies have shown that cells from different age groups or genetic backgrounds can display significant differences in reprogramming efficiency and the quality of the resulting cells. This highlights the need to establish rigorous standards to evaluate in vivo reprogramming efficacy and safety across diverse patient populations. Comprehensive regulatory guidelines are also essential to standardise and coordinate efforts across academic and industry sectors, ensuring responsible and effective development of these technologies.
The future of direct reprogramming lies in continued cross-disciplinary collaborations, integrating insights from molecular biology, bioengineering, and clinical medicine. Advances in single-cell omics technologies are proving invaluable, providing unprecedented insights into the heterogeneous and asynchronous nature of reprogramming, identifying alternative routes, and pinpointing genes and mechanisms that need to be inhibited or activated for precise cell type generation. Combining these mechanistic insights with technical advancements in 3D imaging, tissue reconstruction, and novel biomaterials promises to unlock new therapeutic opportunities, bringing us closer to a future where damaged organs can be repaired or replaced through the remarkable plasticity of our own cells.
Frequently Asked Questions (FAQs)
Q1: What is the main difference between direct reprogramming and iPSC reprogramming?
A1: Direct reprogramming (transdifferentiation) converts one somatic cell type directly into another without passing through a pluripotent stem cell stage. iPSC reprogramming first converts somatic cells into induced pluripotent stem cells (iPSCs), which then differentiate into the desired cell type. Direct reprogramming is generally faster and avoids the risks associated with pluripotency.
Q2: Why is direct reprogramming considered promising for regenerative medicine?
A2: It offers the potential for in situ (on-site) cell conversion within the body, reducing the need for complex ex vivo procedures, large-scale cell expansion, and transplantation. This could simplify treatments, lower costs, and reduce risks like immune rejection.
Q3: What are pioneer factors in the context of cell reprogramming?
A3: Pioneer factors are a special type of transcription factor that can bind to and 'open up' tightly packed chromatin (DNA) regions that are normally inaccessible. This allows other transcription factors to then bind and initiate the gene expression programme for the new cell identity, making them crucial for initiating cell fate change.
Q4: What role do small molecules play in direct reprogramming?
A4: Small molecules can induce or enhance direct reprogramming by altering cellular pathways and gene expression. They offer advantages over viral methods as they are non-immunogenic, cost-effective, and easier to standardise, making them attractive for clinical applications.
Q5: What are some of the biggest challenges facing the clinical translation of direct reprogramming?
A5: Key challenges include improving conversion efficiency and cell maturation, developing safe and precise delivery methods, addressing patient variability (age, genetics), and establishing clear regulatory guidelines for clinical use.
If you want to read more articles similar to Cell Fate: Reprogramming's Profound Impact, you can visit the Automotive category.